Model assumptions and graphical diagnostic tools
Jan Vávra
Exercise 9
Download this R markdown as: R, Rmd.
Outline of this lab session:
- Model assumptions and diagnostic tools
- Graphical diagnostic tools
Loading the data and libraries
Firstly, some necessary R libraries and the working dataset:
library("colorspace") # colors defined by function
library("mffSM") # data and plotLM()
library("splines") # B-splines modelling
library("MASS") # function stdres for standardized residuals
library("fBasics") # dagoTest()
library("car") # ncvTest()
library("lmtest") # bptest()We are going to use the dataset Dris from Exercise
6:
data(Dris, package = "mffSM")
# help("Dris")
head(Dris)## yield N P K Ca Mg
## 1 5.47 470 47 320 47 11
## 2 5.63 530 48 357 60 16
## 3 5.63 530 48 310 63 16
## 4 4.84 482 47 357 47 13
## 5 4.84 506 48 294 52 15
## 6 4.21 500 45 283 61 14summary(Dris)## yield N P K Ca Mg
## Min. :1.920 Min. :268.0 Min. :32.00 Min. :200.0 Min. :29.00 Min. : 8.00
## 1st Qu.:4.040 1st Qu.:427.0 1st Qu.:43.00 1st Qu.:338.0 1st Qu.:41.75 1st Qu.:10.00
## Median :4.840 Median :473.0 Median :49.00 Median :377.0 Median :49.00 Median :11.00
## Mean :4.864 Mean :469.9 Mean :48.65 Mean :375.4 Mean :51.45 Mean :11.64
## 3rd Qu.:5.558 3rd Qu.:518.0 3rd Qu.:54.00 3rd Qu.:407.0 3rd Qu.:59.00 3rd Qu.:13.00
## Max. :8.792 Max. :657.0 Max. :72.00 Max. :580.0 Max. :95.00 Max. :22.00par(mar = c(4,4,0.5,0.5))
plot(yield ~ Mg, data = Dris, lwd=2, col="blue4", bg="skyblue", pch=21)
1. Basic model diagnostics
We will use the following model at the beginning:
aLogMg <- lm(yield ~ log2(Mg), data = Dris)
summary(aLogMg) ##
## Call:
## lm(formula = yield ~ log2(Mg), data = Dris)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.1194 -0.7412 -0.0741 0.7451 3.9841
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 1.4851 0.7790 1.907 0.0574 .
## log2(Mg) 0.9605 0.2208 4.349 1.77e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.07 on 366 degrees of freedom
## Multiple R-squared: 0.04915, Adjusted R-squared: 0.04655
## F-statistic: 18.92 on 1 and 366 DF, p-value: 1.772e-05The assumptions on model error terms \(\varepsilon_i\) could be summarized as follows:
- independence of the observations,
- zero mean, \(\mathsf{E}\, \varepsilon_i = 0, \forall i\),
- homoscedasticity, \(\mathsf{var}\, \varepsilon_i = \sigma^2 \forall i\),
- normality.
It is only natural to inspect the model residuals to judge whether these assumptions are fulfilled. There are two types of residuals:
- Raw residuals \(U_1,
\dots, U_n\), that are supposed to satisfy:
- \(\mathsf{E}\, [U_i \mid \boldsymbol{X}_i] = 0\)
- \(\mathsf{var}\, [U_i \mid \boldsymbol{X}_i] = \sigma^2 m_{ii}\), where \(m_{ii}\) is a diagonal element of the matrix \(\mathbb{M} = (\mathbb{I} - \mathbb{H})\)
- in a normal linear model, it also holds that \(U_i\) follow the normal distribution, however, with different variance for different \(i \in \{1, \dots, n\}\)
- Standardized residuals \(V_i = U_i / \sqrt{\widehat{\sigma}_n \, m_{ii}} =
U_i \sqrt{\frac{n-p}{\mathsf{RSS} \, m_{ii}} }\), for \(i = 1, \dots, n\) that satify
- \(\mathsf{E}\, [V_i \mid \boldsymbol{X}_i] = 0\)
- \(\mathsf{var}\, [V_i \mid \boldsymbol{X}_i] = 1\)
- however, the standardized residuals are not normal – not even under the normal linear model
Here are the three most important nicely formatted graphical diagnostic tools:
par(mar = c(4,4,3,0.5))
plotLM(aLogMg) # function from 'mffSM' -1.png)
Individual work
How can we manually reconstruct the plots above?
- First plot (Residuals vs Fitted):
par(mfrow = c(1,2), mar = c(4,4,2,0.5))
plot(aLogMg, which = 1)
### manual computation
plot(fitted(aLogMg), residuals(aLogMg),
lwd=2, col="blue4", bg="skyblue", pch=21)
abline(h=0, col = "grey", lty = 3)
lines(lowess(fitted(aLogMg), residuals(aLogMg)), col="red", lwd=3)
wmaxabs <- order(abs(residuals(aLogMg)), decreasing = TRUE)[1:3]
text(fitted(aLogMg)[wmaxabs], residuals(aLogMg)[wmaxabs], label = wmaxabs, pos = 4)
- Second plot (QQ plot):
par(mfrow = c(1,2), mar = c(4,4,2,0.5))
plot(aLogMg, which = 2, ask = FALSE)
### standardized residuals
H = qr.Q(aLogMg[["qr"]]) %*% t(qr.Q(aLogMg[["qr"]]))
M = diag(nrow(H))-H
stdres = residuals(aLogMg)/sqrt(deviance(aLogMg)/aLogMg$df.residual*diag(M))
qq = qqnorm(stdres, plot.it=FALSE)
# all.equal(stdres(aLogMg), stdres) # --> TRUE (computed by MASS function)
# all.equal(qq$y, stdres) # --> TRUE (will be on y-axis)
# all.equal(sort(qq$x), qnorm((1:nrow(Dris)-0.5)/nrow(Dris)))
plot(qq$x, qq$y, lwd=2, col="blue4", bg="skyblue", pch=21)
qqline(stdres, lwd=2, lty=3)
text(qq$x[wmaxabs], qq$y[wmaxabs], label = wmaxabs, pos = ifelse(qq$x[wmaxabs] < 0, 4, 2))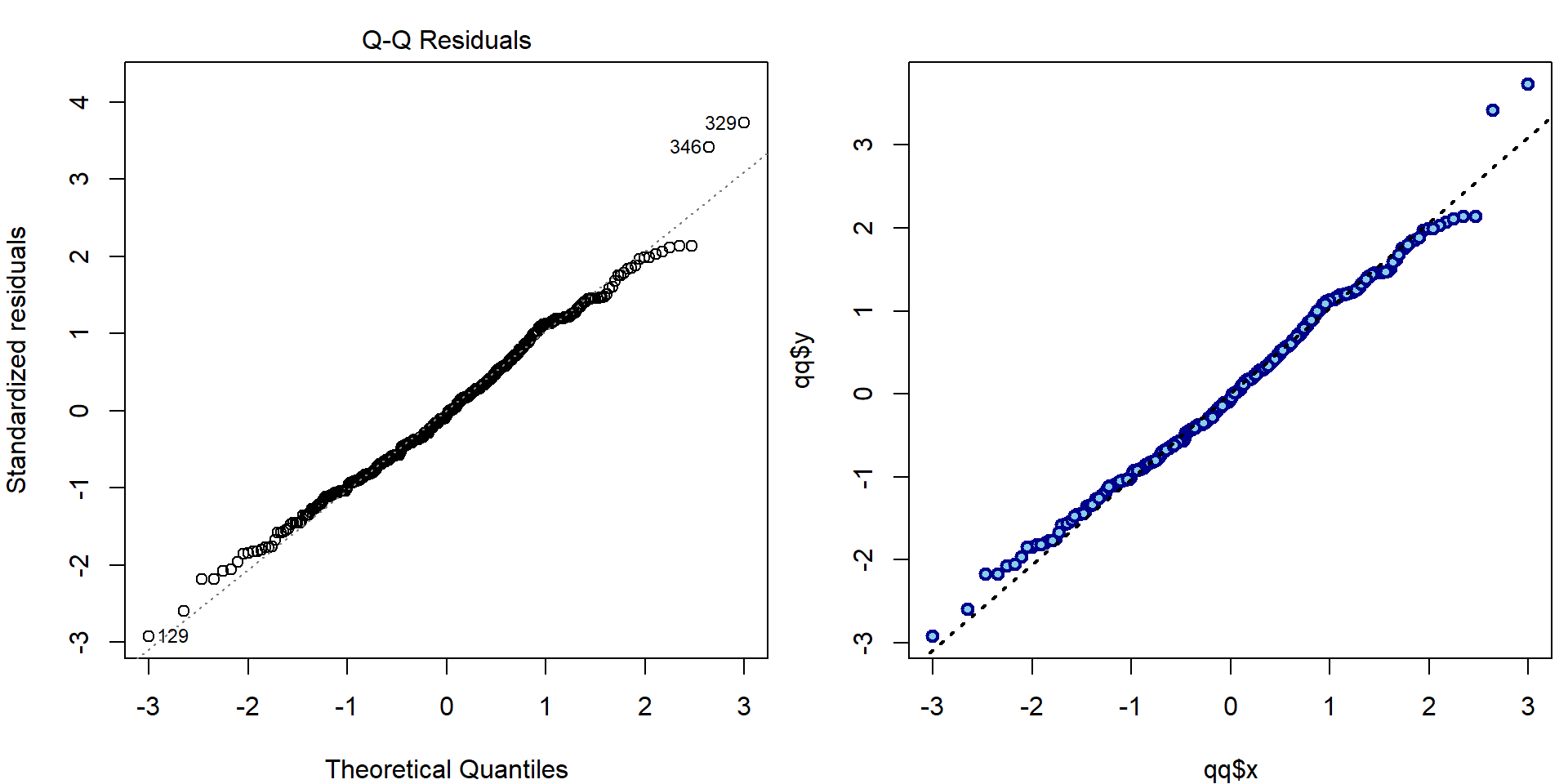
- Third plot (Scale-Location plot)
par(mfrow = c(1,2), mar = c(4,4,2,0.5))
plot(aLogMg, which = 3, ask = FALSE)
### manual computation
plot(fitted(aLogMg), sqrt(abs(stdres)),
lwd=2, col="blue4", bg="skyblue", pch=21)
lines(lowess(fitted(aLogMg), sqrt(abs(stdres))), col="red", lwd=3)
text(fitted(aLogMg)[wmaxabs], sqrt(abs(stdres))[wmaxabs], label = wmaxabs, pos = 4)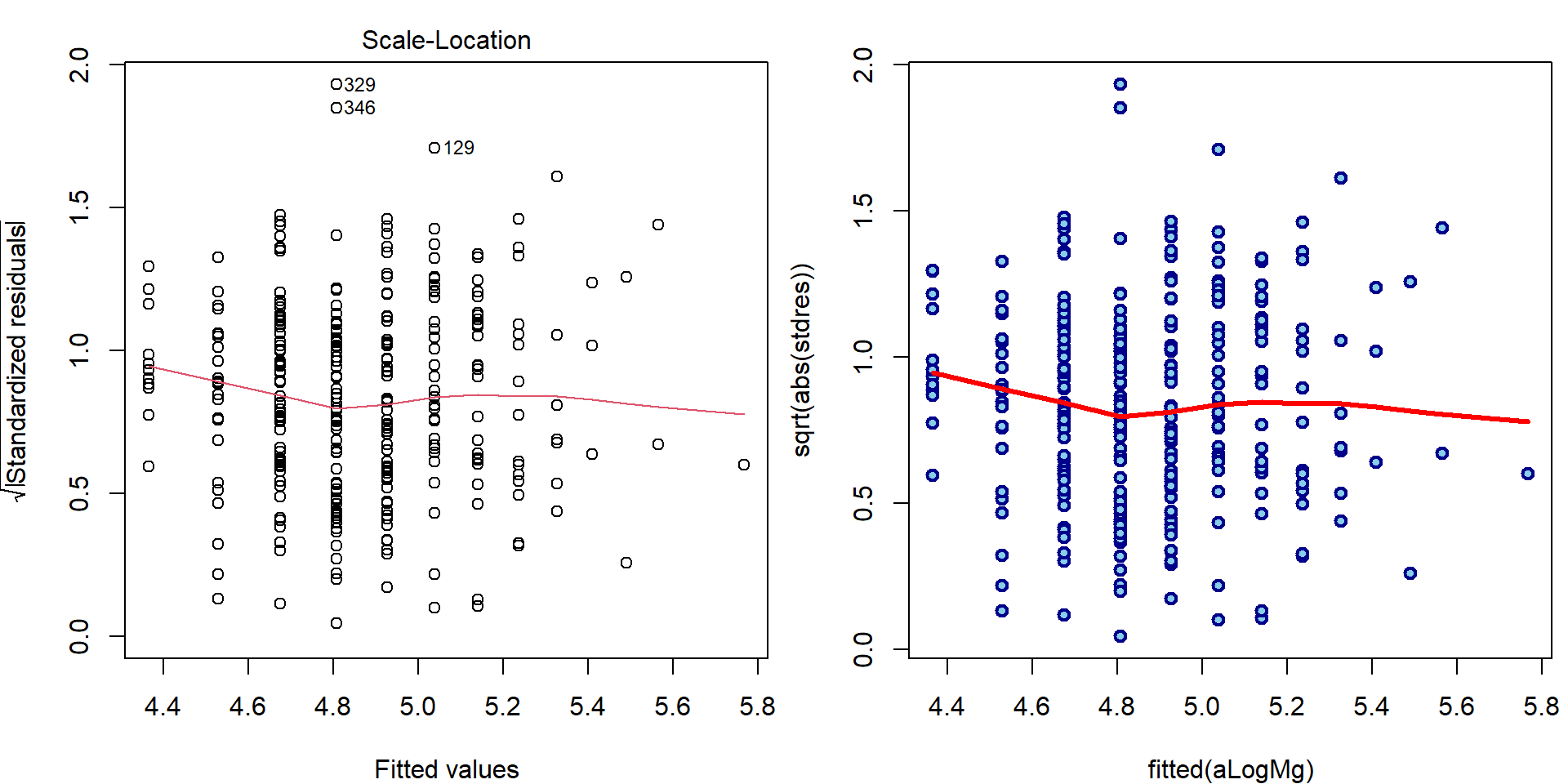
Note, that there are actually infinitely many possibilities how to check that \(\mathsf{E}\,[U_i \mid \boldsymbol{X}_i] = 0\). A scatterplot of fitted values vs. residuals is only one of them. Other common possibility is to plot the covariate values vs. the residuals.
par(mfrow = c(1,2), mar = c(4,4,2,0.5))
plot(log2(Dris$Mg), residuals(aLogMg),
xlab=expression(log[2](Mg)), ylab="Residuals",
main="Residuals vs. covariate in model",
pch = 21, col = "blue4", bg = "skyblue")
lines(lowess(residuals(aLogMg)~log2(Dris$Mg)), col="red")
plot(log2(Dris$Ca), residuals(aLogMg),
xlab=expression(log[2](Ca)), ylab="Residuals",
main="Residuals vs. covariate not in model",
pch = 21, col = "blue4", bg = "skyblue")
lines(lowess(residuals(aLogMg)~log2(Dris$Ca)), col="red")
Also check the following and explain the results:
sum(aLogMg$residuals)## [1] -5.522666e-14sum(aLogMg$residuals*log2(Dris$Mg))## [1] -2.58342e-13Compare the plots for model aLogMg with the interaction
model from previous exercise:
par(mar = c(4,4,3,0.5))
plotLM(lm(yield ~ log2(Mg) + log2(Ca) + I(log2(Mg) * log2(Ca)), data = Dris))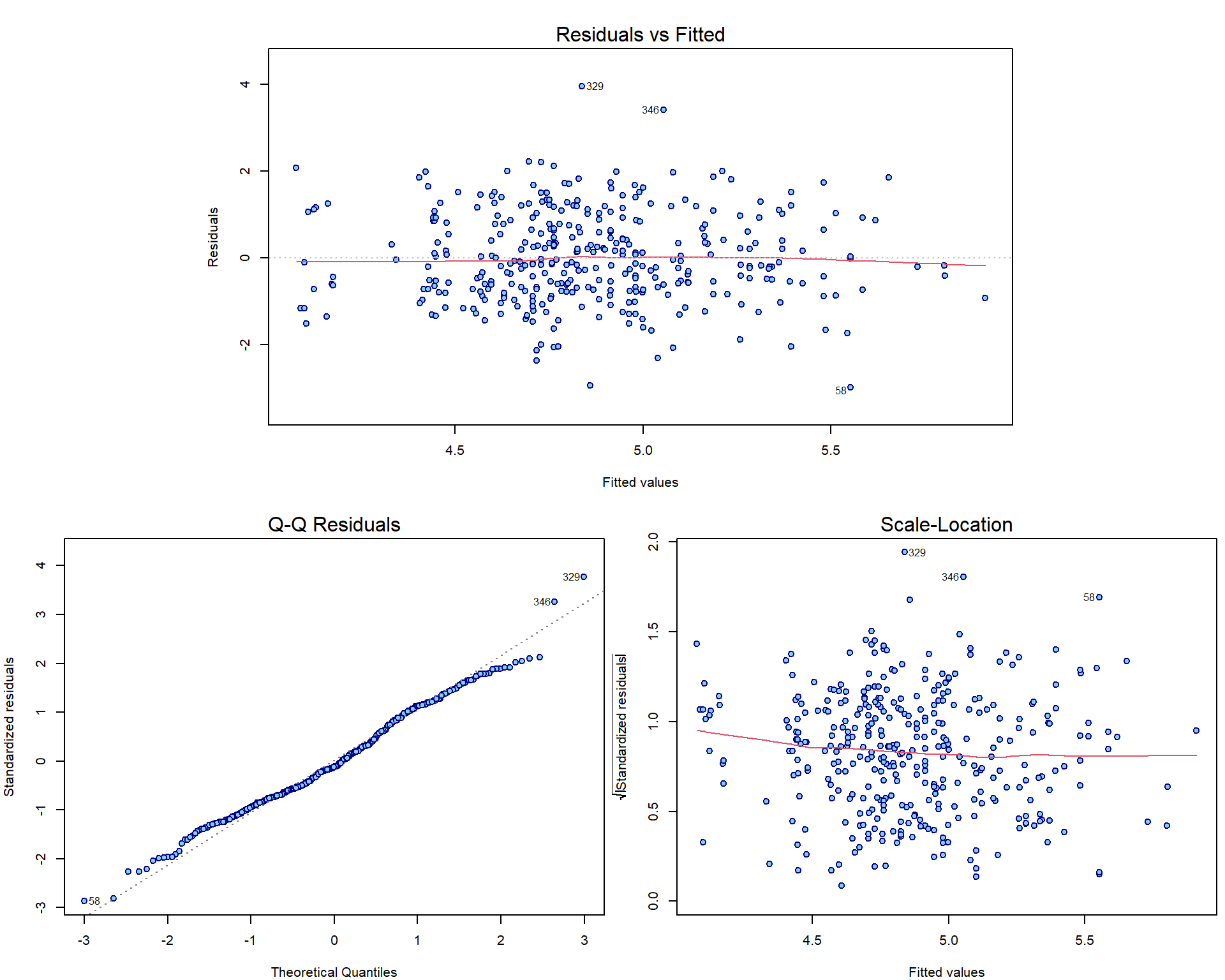
2. Fitting correct \(E[Y \mid \mathbf{X}]\)
The key to make sure \(E[Y \mid \mathbf{X}]\) is fitted correctly is to explore, whether proper parametrizations of covariates are used, which was covered by previous exercises. Here we will demonstrate the necessity of correct parametrization on a data set with extreme behaviour of the response.
A dataset providing a series of (independent) measurements of head
acceleration in a simulated motorcycle accident (used to test crash
helmets) is given in the datafile Motorcycle (within the R
package mffSM).
data(Motorcycle, package = "mffSM")
# help(Motorcycle)
head(Motorcycle)## time haccel
## 1 2.4 0.0
## 2 2.6 -1.3
## 3 3.2 -2.7
## 4 3.6 0.0
## 5 4.0 -2.7
## 6 6.2 -2.7dim(Motorcycle)## [1] 133 2summary(Motorcycle)## time haccel
## Min. : 2.40 Min. :-134.00
## 1st Qu.:15.80 1st Qu.: -54.90
## Median :24.00 Median : -13.30
## Mean :25.55 Mean : -25.55
## 3rd Qu.:35.20 3rd Qu.: 0.00
## Max. :65.60 Max. : 75.00We are primarily interested in how the head acceleration (covariate
haccel) depends on the time after the accident (given in
miliseconds – the time covariate).
par(mfrow = c(1,1), mar = c(4,4,1,1))
plot(haccel ~ time, data = Motorcycle, pch = 21, bg = "orange", col = "red3",
xlab = "Time [ms]", ylab = "Head acceleration [g]")
fit0 = lm(haccel~time, data=Motorcycle)
abline(fit0)
Using the diagnostic plots, we can clearly see that the model denoted
as fit0 is hardly appropriate:
par(mar = c(4,4,3,0.5))
plotLM(fit0)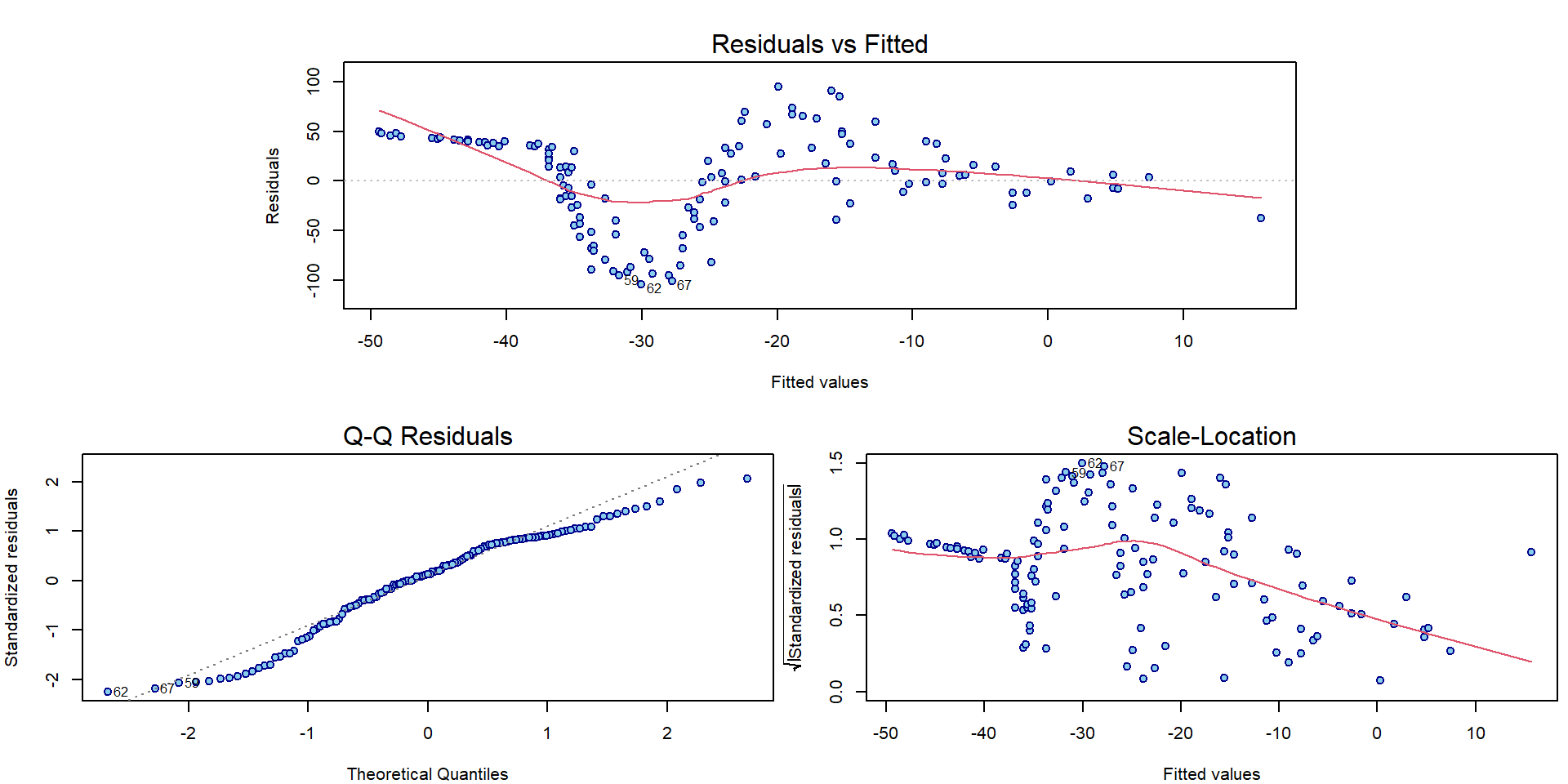
As far as there is a quite flexible shape observed in the plot we need some more flexible model to capture the structure of the underlying conditional expectation of the head acceleration given the time.
Polynomials and splines
We will use polynomials (of different degrees) for fitting the model (while still staying within the linear regression framework).
fit6 <- lm(haccel~poly(time, 6, raw=TRUE), data=Motorcycle)
fit12 <- lm(haccel~poly(time, 12, raw=TRUE), data=Motorcycle)The model can be visualized as follows:
xgrid <- seq(0, 70, length = 500)
par(mfrow = c(1,1), mar = c(4,4,2,1))
plot(haccel ~ time, data = Motorcycle, pch = 21, bg = "orange", col = "red3",
main = "Fitted polynomial curves",
xlab = "Time [ms]", ylab = "Head acceleration [g]")
lines(xgrid, predict(fit6, newdata=data.frame(time=xgrid)), lwd=2, col="blue")
lines(xgrid, predict(fit12, newdata=data.frame(time=xgrid)), lwd=2, col="darkgreen")
legend(37,-70, lty=rep(1, 2), col=c("blue", "darkgreen"),
legend=paste("p=", c(6,12),sep=""), lwd=c(2,2), cex=1.2)
Classical polynomials are, however, not computationally very
effective. Especially for higher order of the polynomial approximation
the solution tends to be unstable. To avoid numerical problems you can
use orthogonal polynomials with raw=FALSE, which stabilizes
the magnitude (and standard errors) of estimated coefficients:
summary(fit6)##
## Call:
## lm(formula = haccel ~ poly(time, 6, raw = TRUE), data = Motorcycle)
##
## Residuals:
## Min 1Q Median 3Q Max
## -76.039 -23.622 -0.497 24.032 75.521
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -2.069e+02 4.349e+01 -4.757 5.29e-06 ***
## poly(time, 6, raw = TRUE)1 9.658e+01 1.558e+01 6.199 7.47e-09 ***
## poly(time, 6, raw = TRUE)2 -1.302e+01 1.865e+00 -6.982 1.49e-10 ***
## poly(time, 6, raw = TRUE)3 7.160e-01 1.012e-01 7.078 9.04e-11 ***
## poly(time, 6, raw = TRUE)4 -1.870e-02 2.732e-03 -6.846 2.97e-10 ***
## poly(time, 6, raw = TRUE)5 2.321e-04 3.572e-05 6.496 1.73e-09 ***
## poly(time, 6, raw = TRUE)6 -1.102e-06 1.799e-07 -6.128 1.05e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 33.09 on 126 degrees of freedom
## Multiple R-squared: 0.5524, Adjusted R-squared: 0.5311
## F-statistic: 25.91 on 6 and 126 DF, p-value: < 2.2e-16summary(lm(haccel~poly(time, 6, raw=FALSE), data=Motorcycle))##
## Call:
## lm(formula = haccel ~ poly(time, 6, raw = FALSE), data = Motorcycle)
##
## Residuals:
## Min 1Q Median 3Q Max
## -76.039 -23.622 -0.497 24.032 75.521
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -25.546 2.869 -8.903 5.01e-15 ***
## poly(time, 6, raw = FALSE)1 160.611 33.091 4.854 3.52e-06 ***
## poly(time, 6, raw = FALSE)2 99.015 33.091 2.992 0.00333 **
## poly(time, 6, raw = FALSE)3 -244.969 33.091 -7.403 1.67e-11 ***
## poly(time, 6, raw = FALSE)4 64.683 33.091 1.955 0.05283 .
## poly(time, 6, raw = FALSE)5 171.291 33.091 5.176 8.68e-07 ***
## poly(time, 6, raw = FALSE)6 -202.776 33.091 -6.128 1.05e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 33.09 on 126 degrees of freedom
## Multiple R-squared: 0.5524, Adjusted R-squared: 0.5311
## F-statistic: 25.91 on 6 and 126 DF, p-value: < 2.2e-16Another possible solution how to introduce some reasonable stability into the model where some higher order polynomials are used, are splines.
### Choice of knots
knots <- c(0, 11, 12, 13, 20, 30, 32, 34, 40, 50, 70)
(inner <- knots[-c(1, length(knots))]) # inner knots ## [1] 11 12 13 20 30 32 34 40 50(bound <- knots[c(1, length(knots))]) # boundary knots ## [1] 0 70### B-splines of degree 3
DEGREE <- 3 # piecewise cubic
Bgrid <- bs(xgrid, knots = inner, Boundary.knots = bound, degree = DEGREE,
intercept = TRUE)
dim(Bgrid) # 13-dimensional linear space of piecewise polynomials## [1] 500 13The B-spline basis is a set of the following functions:
COL <- rainbow_hcl(ncol(Bgrid), c = 80)
par(mfrow = c(1,1), mar = c(4,4,1,1))
plot(xgrid, Bgrid[,1], type = "l", col = COL[1], lwd = 2, xlab = "x",
ylab = "B(x)", xlim = range(knots), ylim = range(Bgrid, na.rm = TRUE))
for (j in 2:ncol(Bgrid)) lines(xgrid, Bgrid[,j], col = COL[j], lwd = 2)
points(knots, rep(0, length(knots)),
pch = 21, bg = "blue", col = "skyblue", cex = 1.5)
Thus, we can use B-splines to fit the model.
More details about the B-splines can be found, for instance, here.
m1 <- lm(haccel ~ bs(time, knots = inner, Boundary.knots = bound,
degree = DEGREE, intercept = TRUE) - 1, data = Motorcycle)This is the fitted curve:
par(mfrow = c(1,1), mar = c(4,4,2,1))
plot(haccel ~ time, data = Motorcycle, pch = 21, bg = "orange", col = "red3",
main = "Fitted B-spline curve",
xlab = "Time [ms]", ylab = "Head acceleration [g]")
lines(xgrid, predict(m1, newdata=data.frame(time=xgrid)), lwd=2, col="blue")
names(m1$coefficients) <- paste("B", 1:ncol(Bgrid))
summary(m1)##
## Call:
## lm(formula = haccel ~ bs(time, knots = inner, Boundary.knots = bound,
## degree = DEGREE, intercept = TRUE) - 1, data = Motorcycle)
##
## Residuals:
## Min 1Q Median 3Q Max
## -71.857 -11.740 -0.268 10.356 55.249
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## B 1 -11.614 83.252 -0.140 0.889
## B 2 12.450 81.282 0.153 0.879
## B 3 -13.988 45.345 -0.308 0.758
## B 4 2.984 18.894 0.158 0.875
## B 5 6.115 12.139 0.504 0.615
## B 6 -237.283 18.850 -12.588 < 2e-16 ***
## B 7 17.346 14.094 1.231 0.221
## B 8 53.248 12.635 4.214 4.88e-05 ***
## B 9 5.102 13.051 0.391 0.697
## B 10 12.609 17.146 0.735 0.464
## B 11 -34.949 38.161 -0.916 0.362
## B 12 58.255 65.045 0.896 0.372
## B 13 -93.068 75.924 -1.226 0.223
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 22.61 on 120 degrees of freedom
## Multiple R-squared: 0.8447, Adjusted R-squared: 0.8278
## F-statistic: 50.19 on 13 and 120 DF, p-value: < 2.2e-16Since the intercept is included within the B-spline basis
(intercept = TRUE in bs), we omitted it in the
formula with -1. However, this use comes with some issues
deeply rooted within the implementation. For example, neither
Multiple R-squared nor the F-statistic
quantity in the output correspond to correct numbers. Simply put,
summary.lm thinks the intercept is not included despite
still being a linear combination of given columns.
Individual work
Can you explain why the values reported for \(R^2\) and \(R_{adj}^2\) are not appropriate? What is the right meaning of these numbers? Can you manually reconstruct them?
The \(R^2\) value from the model is
summary(m1)$r.squared## [1] 0.8446597and the manual calculations will give
1 - deviance(m1)/sum(Motorcycle$haccel^2)## [1] 0.8446597sum(fitted(m1)^2)/sum(Motorcycle$haccel^2)## [1] 0.8446597Interpretation of the F-test from the summary output above is a test of a submodel
summary(m1)$fstatistic## value numdf dendf
## 50.19211 13.00000 120.00000# the usual F-test (correct)
m2 <- lm(haccel ~ 1, data = Motorcycle)
anova(m2, m1)## Analysis of Variance Table
##
## Model 1: haccel ~ 1
## Model 2: haccel ~ bs(time, knots = inner, Boundary.knots = bound, degree = DEGREE,
## intercept = TRUE) - 1
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 132 308223
## 2 120 61362 12 246861 40.23 < 2.2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# the wrong F-test (in summary m1)
(m0 <- lm(haccel ~ -1, data=Motorcycle))##
## Call:
## lm(formula = haccel ~ -1, data = Motorcycle)
##
## No coefficientsanova(m0 ,m1)## Analysis of Variance Table
##
## Model 1: haccel ~ -1
## Model 2: haccel ~ bs(time, knots = inner, Boundary.knots = bound, degree = DEGREE,
## intercept = TRUE) - 1
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 133 395017
## 2 120 61362 13 333655 50.192 < 2.2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Nevertheless, as the intercept parameter is in the linear span of the columns of \(\mathbb{X}\), it makes sense to calculate the (usual) multiple \(R^2\) value. This can be done ‘manually’ as follows:
var(fitted(m1)) / var(Motorcycle$haccel)## [1] 0.8009163cor(Motorcycle$haccel, fitted(m1))^2## [1] 0.80091631 - deviance(m1)/deviance(m2)## [1] 0.8009163Alternatively, one can use B-splines (the basis without intercept) with an intercept in the model to get some meaningful numbers in the output:
m3 <- lm(haccel ~ bs(time, knots = inner, Boundary.knots = bound,
degree = DEGREE, intercept = FALSE), data = Motorcycle)
names(m3$coefficients) <- c("(Intercept)", paste("B", 1:12))
summary(m3)##
## Call:
## lm(formula = haccel ~ bs(time, knots = inner, Boundary.knots = bound,
## degree = DEGREE, intercept = FALSE), data = Motorcycle)
##
## Residuals:
## Min 1Q Median 3Q Max
## -71.857 -11.740 -0.268 10.356 55.249
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -11.614 83.252 -0.140 0.8893
## B 1 24.064 161.970 0.149 0.8821
## B 2 -2.374 57.835 -0.041 0.9673
## B 3 14.598 93.327 0.156 0.8760
## B 4 17.729 80.496 0.220 0.8261
## B 5 -225.669 88.698 -2.544 0.0122 *
## B 6 28.960 82.780 0.350 0.7271
## B 7 64.862 84.773 0.765 0.4457
## B 8 16.716 83.997 0.199 0.8426
## B 9 24.223 85.172 0.284 0.7766
## B 10 -23.335 91.337 -0.255 0.7988
## B 11 69.869 105.918 0.660 0.5107
## B 12 -81.454 112.465 -0.724 0.4703
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 22.61 on 120 degrees of freedom
## Multiple R-squared: 0.8009, Adjusted R-squared: 0.781
## F-statistic: 40.23 on 12 and 120 DF, p-value: < 2.2e-16anova(m2,m3)## Analysis of Variance Table
##
## Model 1: haccel ~ 1
## Model 2: haccel ~ bs(time, knots = inner, Boundary.knots = bound, degree = DEGREE,
## intercept = FALSE)
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 132 308223
## 2 120 61362 12 246861 40.23 < 2.2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1What about the interpretation of the classical diagnostic plots for the model fitted with B-splines?
par(mar = c(4,4,3,0.5))
plotLM(m1)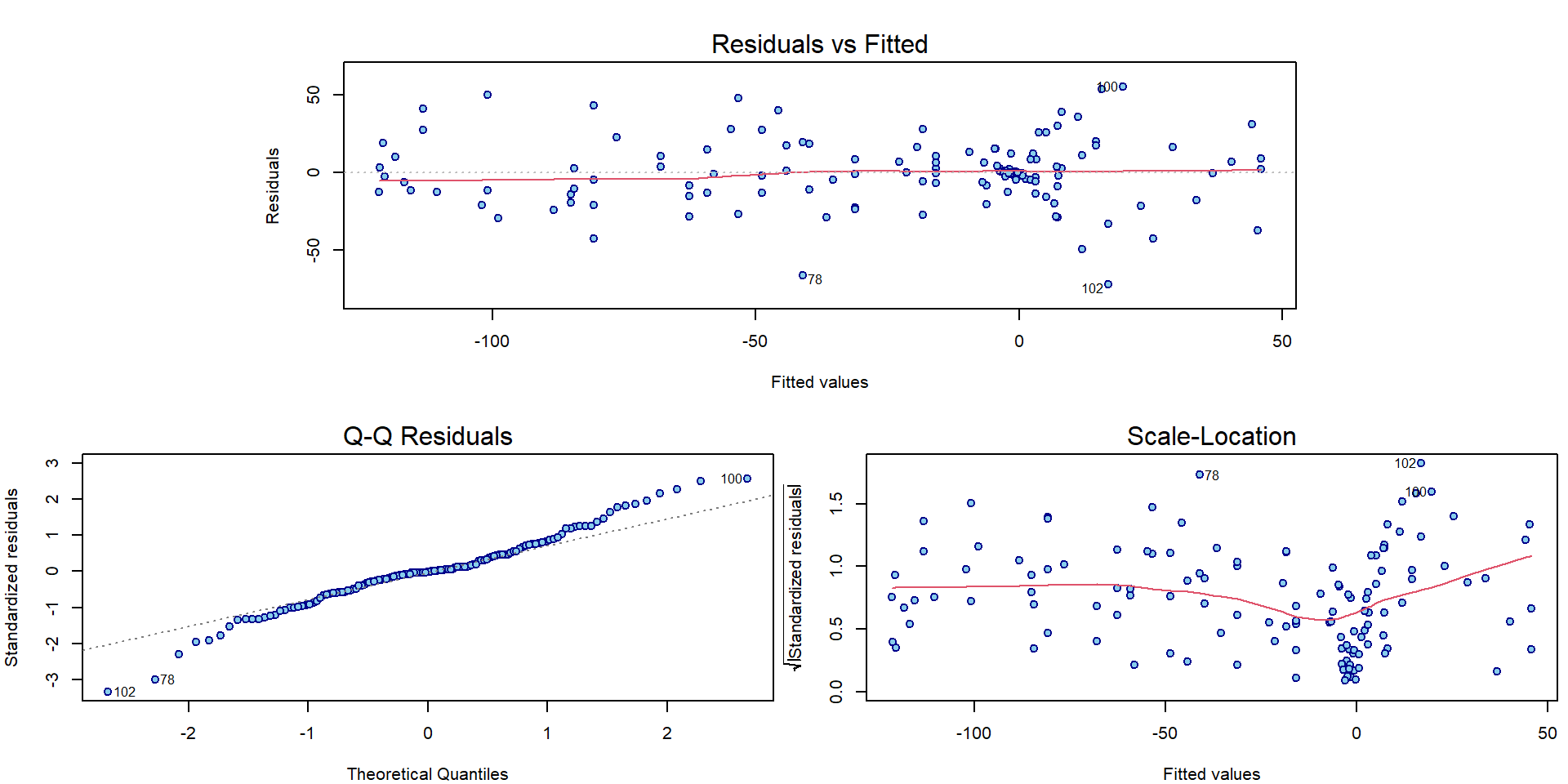
It seems there is some issue with homoscedasticity assumption. Let’s change the x-axis of the third Scale-Location plot to determine how the variability changes:
par(mfrow = c(1,2), mar = c(4,4,2,1))
plot(abs(fitted(m1)), sqrt(abs(stdres(m1))),
main = "Abs(fitted)",
lwd=2, col="blue4", bg="skyblue", pch=21)
lines(lowess(abs(fitted(m1)), sqrt(abs(stdres(m1)))), col="red", lwd=3)
plot(Motorcycle$time, sqrt(abs(stdres(m1))),
main = "Time",
lwd=2, col="blue4", bg="skyblue", pch=21)
lines(lowess(Motorcycle$time, sqrt(abs(stdres(m1)))), col="red", lwd=3)
How to adjust the inference for heteroscedastic models will be shown next exercise.
3. Log transformation of the response
We will use a small dataset (47 observations and 8 different covariates) containing some pieces of information about fires that occurred in Chicago in 1970 while distinguishing for some information provided in the data. Particularly, we will focus on the locality in Chicago (north vs. south) and the proportion of minority citizens. The dataset is loaded from the website and some brief insight is provided below.
chicago <- read.csv("http://www.karlin.mff.cuni.cz/~vavraj/nmfm334/data/chicago.csv",
header=T, stringsAsFactors = TRUE)
chicago <- transform(chicago,
fside = factor(side, levels = 0:1, labels=c("North", "South")))
head(chicago)## minor fire theft old insur income side fside
## 1 10.0 6.2 29 60.4 0.0 11.744 0 North
## 2 22.2 9.5 44 76.5 0.1 9.323 0 North
## 3 19.6 10.5 36 73.5 1.2 9.948 0 North
## 4 17.3 7.7 37 66.9 0.5 10.656 0 North
## 5 24.5 8.6 53 81.4 0.7 9.730 0 North
## 6 54.0 34.1 68 52.6 0.3 8.231 0 Northsummary(chicago)## minor fire theft old insur income
## Min. : 1.00 Min. : 2.00 Min. : 3.00 Min. : 2.00 Min. :0.0000 Min. : 5.583
## 1st Qu.: 3.75 1st Qu.: 5.65 1st Qu.: 22.00 1st Qu.:48.60 1st Qu.:0.0000 1st Qu.: 8.447
## Median :24.50 Median :10.40 Median : 29.00 Median :65.00 Median :0.4000 Median :10.694
## Mean :34.99 Mean :12.28 Mean : 32.36 Mean :60.33 Mean :0.6149 Mean :10.696
## 3rd Qu.:57.65 3rd Qu.:16.05 3rd Qu.: 38.00 3rd Qu.:77.30 3rd Qu.:0.9000 3rd Qu.:11.989
## Max. :99.70 Max. :39.70 Max. :147.00 Max. :90.10 Max. :2.2000 Max. :21.480
## side fside
## Min. :0.0000 North:25
## 1st Qu.:0.0000 South:22
## Median :0.0000
## Mean :0.4681
## 3rd Qu.:1.0000
## Max. :1.0000The data contain information from Chicago insurance redlining in 47 districts in 1970 where
minorstands for the percentage of minority (0 - 100) in a given district,firestands for the number of fires per 100 households during the reference period,theftdenotes the number of reported thefts per 1000 inhabitants,oldgives percentage of residential units built before 1939,insurstates the number of policies per 100 household,incomeis the median income (USD 1000) per household,sidegives the location of the district within Chicago (0 = North, 1 = South).
Exploration
We wish to describe the dependence of the number of fires
(fire) on the percentage of minority (minor).
We are also aware of the fact that the form of this dependence may
differ in north and south parts of Chicago.
par(mfrow = c(1,1), mar = c(4,4,0.5,0.5))
XLAB <- "Minority (%)"
YLAB <- "Fires (#/100 households)"
COLS <- c(North = "darkgreen", South = "red")
BGS <- c(North = "aquamarine", South = "orange")
PCHS <- c(North = 21, South = 23)
plot(fire ~ minor, data = chicago,
xlab = XLAB, ylab = YLAB,
col = COLS[fside], bg = BGS[fside], pch = PCHS[fside])
legend("topleft", legend = c("North", "South"), bty = "n",
pch = PCHS, col = COLS, pt.bg = BGS)
lines(with(subset(chicago, fside == "North"), lowess(minor, fire)),
col = COLS["North"], lwd = 2)
lines(with(subset(chicago, fside == "South"), lowess(minor, fire)),
col = COLS["South"], lwd = 2)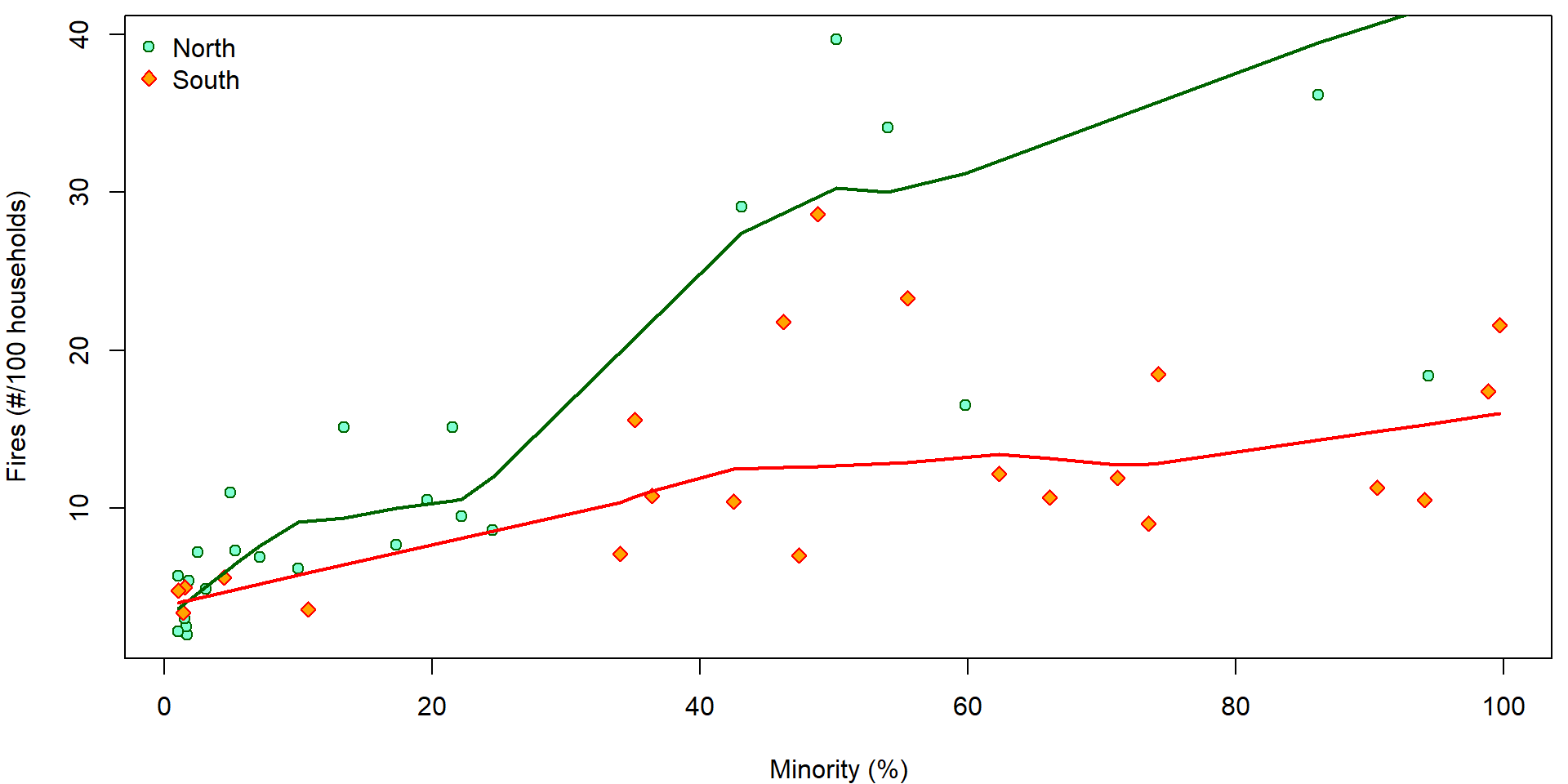
Lets fit different lines for the two sides of Chicago.
mlines <- lm(fire ~ minor * fside, data = chicago)
summary(mlines)##
## Call:
## lm(formula = fire ~ minor * fside, data = chicago)
##
## Residuals:
## Min 1Q Median 3Q Max
## -17.262 -3.340 -1.474 3.075 18.303
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 5.19515 1.69212 3.070 0.0037 **
## minor 0.32274 0.04896 6.592 5.03e-08 ***
## fsideSouth 1.37454 3.10252 0.443 0.6600
## minor:fsideSouth -0.20812 0.06589 -3.159 0.0029 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 6.535 on 43 degrees of freedom
## Multiple R-squared: 0.5387, Adjusted R-squared: 0.5065
## F-statistic: 16.74 on 3 and 43 DF, p-value: 2.376e-07And create the corresponding diagnostic plots:
par(mar = c(4,4,2,0.5))
plotLM(mlines)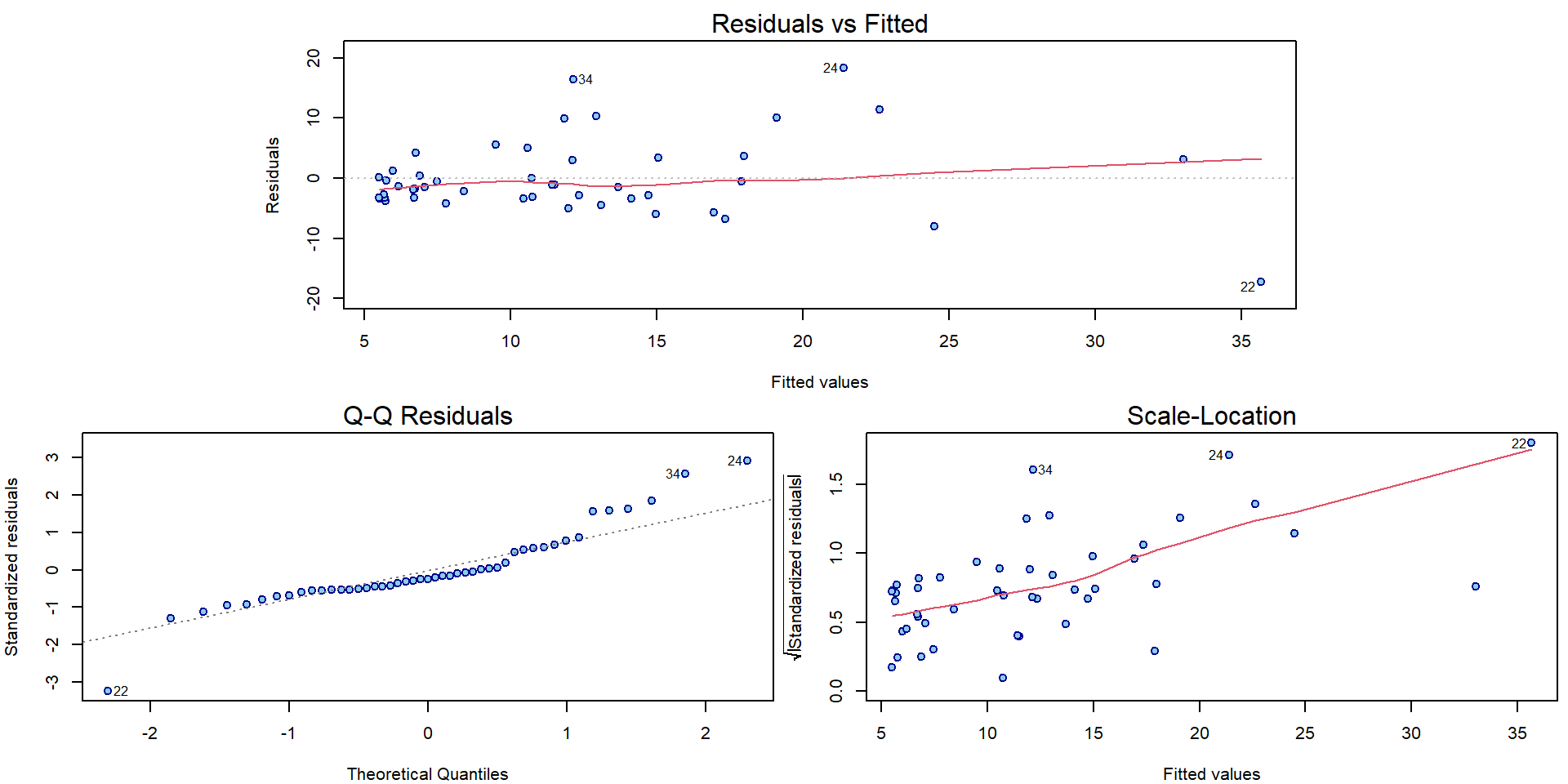
We observe some issues with normality assumption, but especially large issues with increasing variability.
Some statistical tests for normality applied to residuals that are heteroscedastic (even in homoscedastic model):
shapiro.test(residuals(mlines)) ##
## Shapiro-Wilk normality test
##
## data: residuals(mlines)
## W = 0.90361, p-value = 0.0009409dagoTest(residuals(mlines))##
## Title:
## D'Agostino Normality Test
##
## Test Results:
## STATISTIC:
## Chi2 | Omnibus: 9.4764
## Z3 | Skewness: 2.1817
## Z4 | Kurtosis: 2.1717
## P VALUE:
## Omnibus Test: 0.008754
## Skewness Test: 0.02913
## Kurtosis Test: 0.02987Some statistical tests for normality applied to standardized residuals that are not normal (even in normal linear model):
shapiro.test(stdres(mlines)) ##
## Shapiro-Wilk normality test
##
## data: stdres(mlines)
## W = 0.90005, p-value = 0.0007228dagoTest(stdres(mlines))##
## Title:
## D'Agostino Normality Test
##
## Test Results:
## STATISTIC:
## Chi2 | Omnibus: 7.9183
## Z3 | Skewness: 1.3467
## Z4 | Kurtosis: 2.4707
## P VALUE:
## Omnibus Test: 0.01908
## Skewness Test: 0.1781
## Kurtosis Test: 0.01348Statistical test for homoscedasticity (Breusch-Pagan test and Koenker’s studentized version) with alternatives \(\mathsf{var} \left[ Y_i | \mathbf{X}_i\right] = \sigma^2 \exp\left\{\tau \mathsf{E} \left[Y_i | \mathbf{X}_i\right]\right\}\) or in general \(\mathsf{var} \left[ Y_i | \mathbf{X}_i, \mathbf{Z_i}\right] = \sigma^2 \exp\left\{\mathbf{Z}_i^\top \boldsymbol{\tau}\right\}\):
ncvTest(mlines) # default is fitted## Non-constant Variance Score Test
## Variance formula: ~ fitted.values
## Chisquare = 25.2439, Df = 1, p = 5.0519e-07ncvTest(mlines, var.formula = ~fitted(mlines)) # the same as before ## Non-constant Variance Score Test
## Variance formula: ~ fitted(mlines)
## Chisquare = 25.2439, Df = 1, p = 5.0519e-07bptest(mlines, varformula = ~fitted(mlines)) # Koenker's studentized version (robust to non-normality)##
## studentized Breusch-Pagan test
##
## data: mlines
## BP = 13.692, df = 1, p-value = 0.0002153bptest(mlines) # default is the same as model formula##
## studentized Breusch-Pagan test
##
## data: mlines
## BP = 15.212, df = 3, p-value = 0.001644In addition, there is also a bunch of points scattered around the origin and relatively only a few observations are given for larger proportion of the minority or the larger amount of fires. This suggests that log transformation (of the response and the predictor as well) could help.
Interpretation of a model with log-transformed response
Typically, when the residual variance increases with the response expectation, log-transformation of the response often ensures a homoscedastic model (Assumption A3a from the lecture).
Firstly, compare the plot below (on the log scale on \(y\) axes) with the previous plot on the original scale on the \(y\) axes:
par(mfrow = c(1,1), mar = c(4,4,0.5,0.5))
LYLAB <- "Log(Fires)"
plot(log(fire) ~ minor, data = chicago,
xlab = XLAB, ylab = LYLAB,
col = COLS[fside], bg = BGS[fside], pch = PCHS[fside])
legend("topleft", legend = c("North", "South"), bty = "n",
pch = PCHS, col = COLS, pt.bg = BGS)
lines(with(subset(chicago, fside == "North"), lowess(minor, log(fire))),
col = COLS["North"], lwd = 2)
lines(with(subset(chicago, fside == "South"), lowess(minor, log(fire))),
col = COLS["South"], lwd = 2)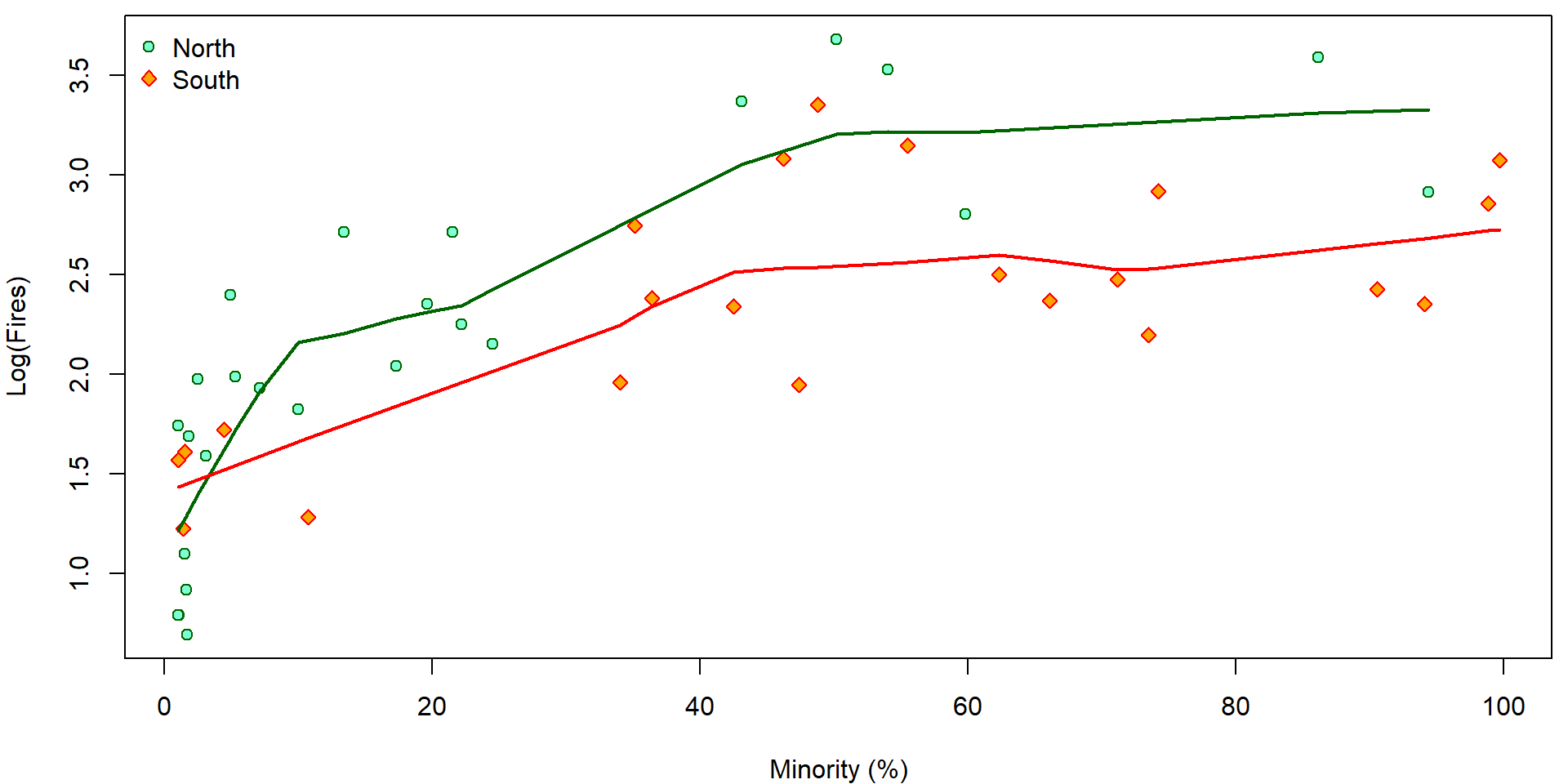
Using the proposed transformation of the response we also change the underlying data. This means, that instead of the model \[ \mathsf{E} [Y_i | \boldsymbol{X}_i] = \boldsymbol{X}_i^\top \boldsymbol{\beta} \] fitted based on the random sample \(\{(Y_i, \boldsymbol{X}_i);\; i = 1, \dots, n\}\) we fit another model of the form \[ \mathsf{E} [\log(Y_i) | \boldsymbol{X}_i] = \boldsymbol{X}_i^\top \widetilde{\boldsymbol{\beta}} \] which is, however, based on the different random sample \(\{(\log(Y_i), \boldsymbol{X}_i);\;i = 1, \dots, n\}\). This has some crucial consequences. The logarithmic transformation will help to deal with some issues regarding heteroscedastic data but, on the other hand, it will introduce new problems regarding the model interpretability.
The idea is that the final model should be (always) interpreted with
respect to the original data - the information about
the number of fires (fire) - not the logarithm of the
number of fires (log(fire)).
For some better interpretation we will also use the log
transformation of the independent variable – the information about the
proportion of minority – minor. What is the interpretation
of the following model:
summary(m <- lm(log(fire) ~ log2(minor) * fside, data = chicago))##
## Call:
## lm(formula = log(fire) ~ log2(minor) * fside, data = chicago)
##
## Residuals:
## Min 1Q Median 3Q Max
## -0.75178 -0.33949 -0.07901 0.34564 0.84890
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 1.01616 0.14635 6.943 1.55e-08 ***
## log2(minor) 0.35961 0.03853 9.332 6.74e-12 ***
## fsideSouth 0.27963 0.26983 1.036 0.3058
## log2(minor):fsideSouth -0.14411 0.05772 -2.497 0.0164 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.4149 on 43 degrees of freedom
## Multiple R-squared: 0.7278, Adjusted R-squared: 0.7088
## F-statistic: 38.33 on 3 and 43 DF, p-value: 3.234e-12The diagnostic plots have improved in terms of mean and variance:
par(mar = c(4,4,2,0.5))
plotLM(m)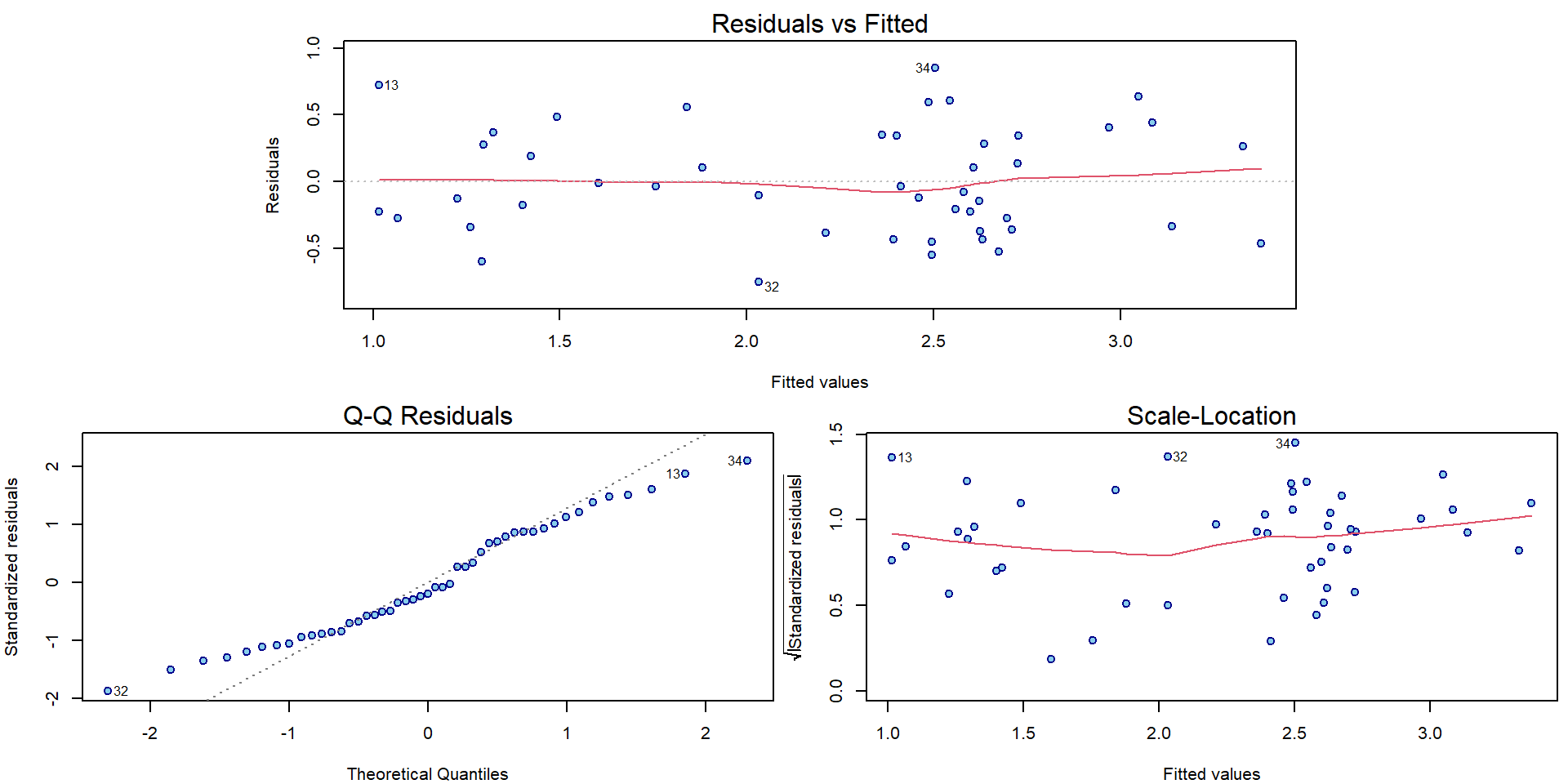
Recall the Jensen inequality and the fact that \[ \mathsf{E} [\log(Y_i) | \boldsymbol{X}_i] \neq \log(\mathsf{E} [Y_i | \boldsymbol{X}_i]) \qquad \text{but rather} \qquad \mathsf{E} [\log(Y_i) | \boldsymbol{X}_i] < \log(\mathsf{E} [Y_i | \boldsymbol{X}_i]). \] And now compare the two models: \[ \begin{aligned} Y_i &= \mathbf{X}_i^\top \boldsymbol{\beta} + \varepsilon_i, \\ \log Y_i = \mathbf{X}_i^\top \widetilde{\boldsymbol{\beta}} + \widetilde{\varepsilon}_i \quad\Longleftrightarrow\quad Y_i &= \exp\left\{\mathbf{X}_i^\top \widetilde{\boldsymbol{\beta}}\right\} \cdot \exp\left\{ \widetilde{\varepsilon}_i \right\}. \end{aligned} \]
What are the consequences with respect to the interpretation of the parameter estimates provided in the summary output above? What would be a suitable interpretation that we can use in this situation?
\[ \dfrac{ \mathsf{E} \left[Y_i | \mathbf{X}_i = \mathbf{x} + \mathbf{e}_j\right] }{ \mathsf{E} \left[Y_i | \mathbf{X}_i = \mathbf{x}\right] } = \dfrac{ \exp \left\{ (\mathbf{x} + \mathbf{e}_j)^\top \widetilde{\boldsymbol{\beta}} \right\} \mathsf{E} \exp\left\{ \widetilde{\varepsilon}_i \right\} }{ \exp \left\{ \mathbf{x}^\top \widetilde{\boldsymbol{\beta}} \right\} \mathsf{E} \exp\left\{ \widetilde{\varepsilon}_i \right\} } = \exp \widetilde{\beta}_j \]
It is easy to visualize the fitted regression function. In addition, we can also add the confidence bands AROUND the regression function and the prediction bands. However, given the previous problems related to the Jensen inequality, only the prediction bands make sense for this model and the confidence (AROUND) bands do not have any good interpretation here.
lpred <- 500
pminor <- seq(1, 100, length = lpred)
pdata <- data.frame(minor = rep(pminor, 2),
fside = factor(rep(c("North", "South"),
each = lpred)))
cifit <- predict(m, newdata = pdata, interval = "confidence", se.fit = TRUE)
pfit <- predict(m, newdata = pdata, interval = "prediction")
YLIM <- range(log(chicago[, "fire"]), pfit)
XLIM <- range(chicago[, "minor"])
par(mfrow = c(1, 2), mar=c(4,4,2,0.5))
for (ss in levels(chicago[, "fside"])){
plot(log(fire) ~ minor, data = subset(chicago, fside == ss), main = ss,
xlab = XLAB, ylab = LYLAB, xlim = XLIM, ylim = YLIM,
col = COLS[fside], bg = BGS[fside], pch = PCHS[fside])
ind <- if (ss == "North") 1:lpred else (lpred + 1):(2 * lpred)
lines(pminor, cifit$fit[ind, "fit"], col = COLS[ss], lwd = 3)
lines(pminor, cifit$fit[ind, "lwr"], col = COLS[ss], lwd = 2, lty = 2)
lines(pminor, cifit$fit[ind, "upr"], col = COLS[ss], lwd = 2, lty = 2)
lines(pminor, pfit[ind, "lwr"], col = COLS[ss], lwd = 2, lty = 3)
lines(pminor, pfit[ind, "upr"], col = COLS[ss], lwd = 2, lty = 3)
legend(x=25, y=1, bty = "n",
legend=c("estim. E[log(Y)|X=x]",
"conf. band around E[log(Y)|X=x]",
"prediction band for log(Y)"),
lwd=c(3,2,2), lty=c(1,2,3), col=COLS[ss])
} 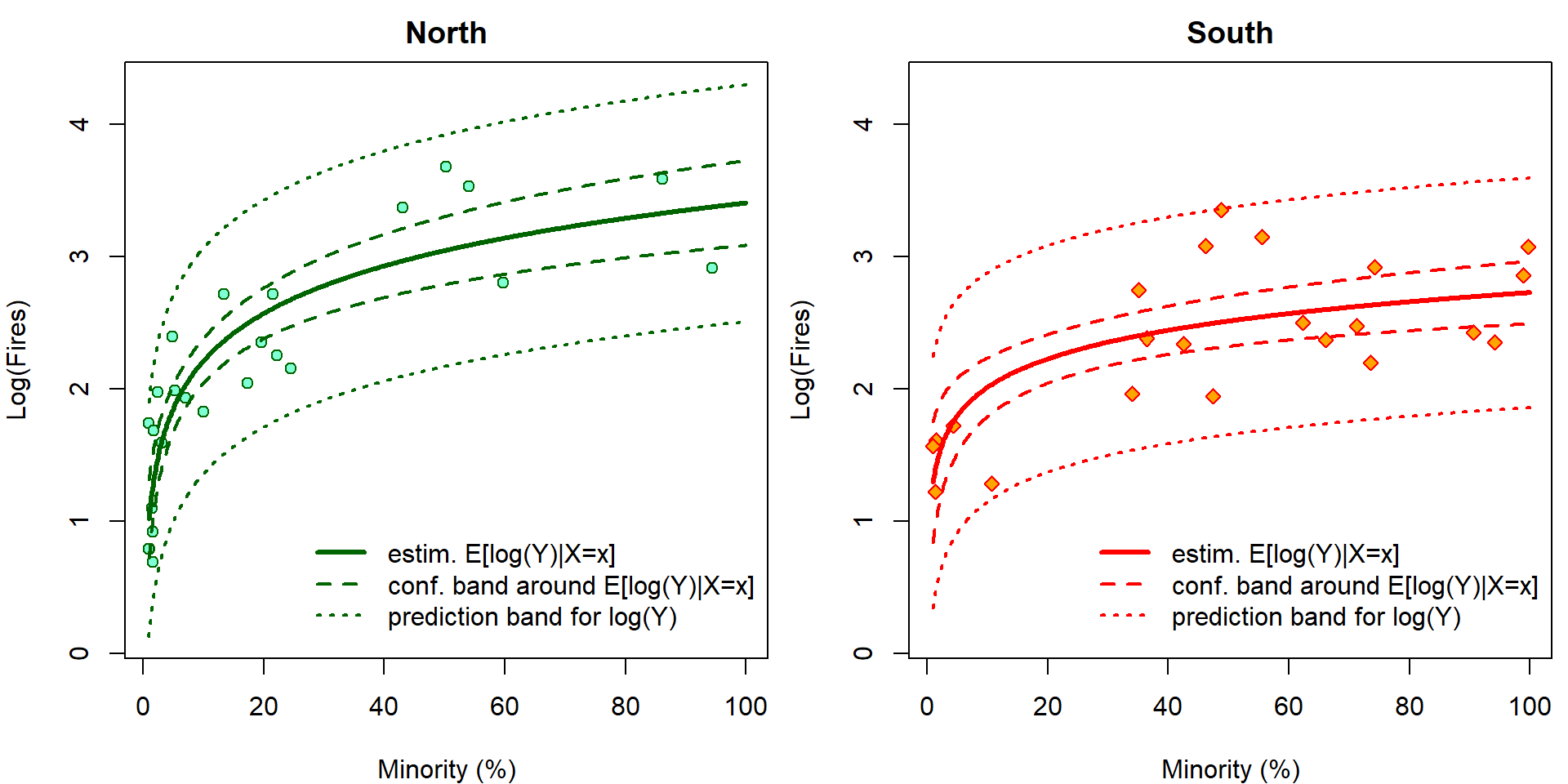
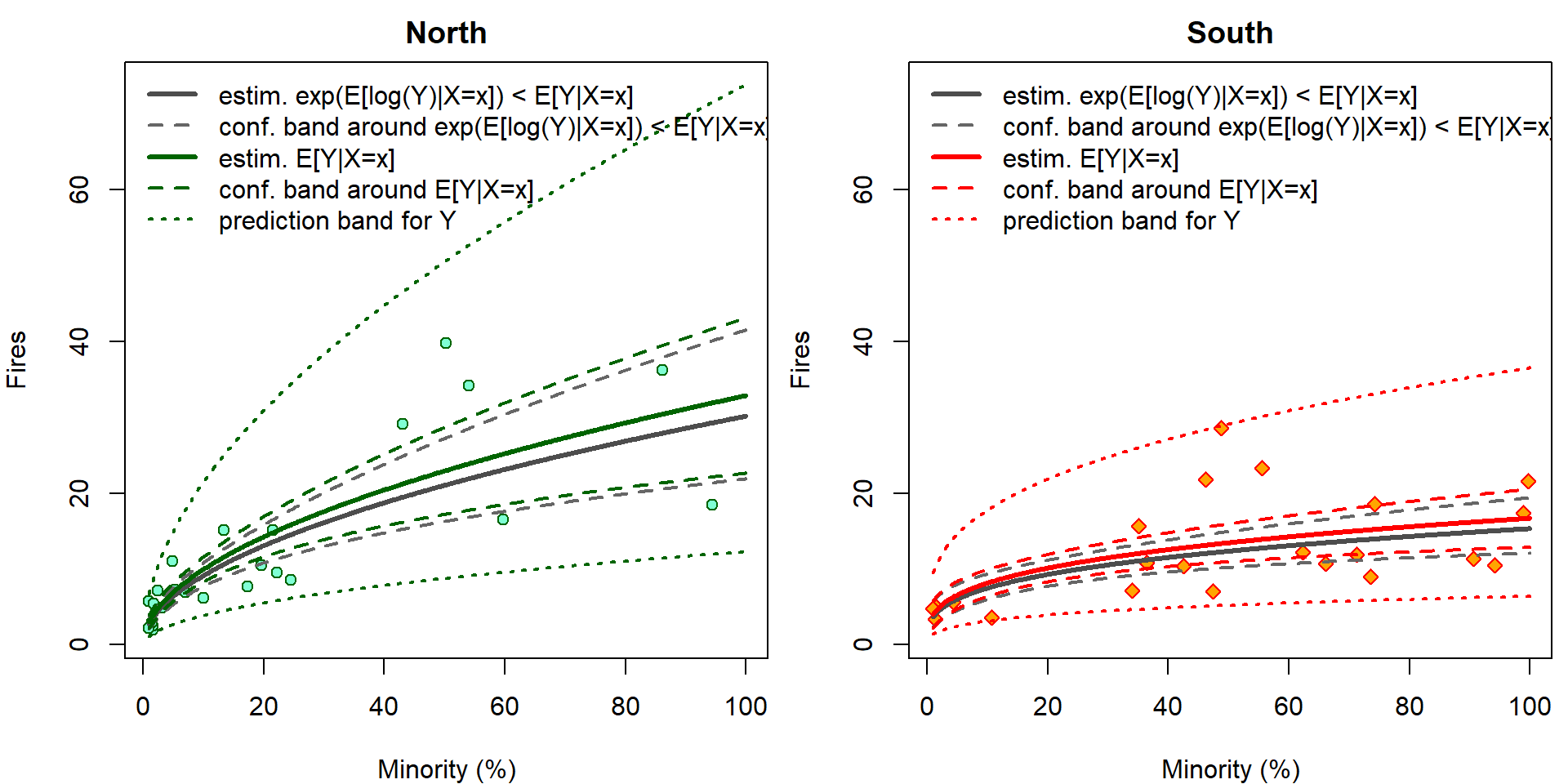
The confidence intervals \[ \mathsf{P} \left( (LCI, UCI)\ni \mathbf{x}^\top \widetilde{\boldsymbol{\beta}} = \mathsf{E}\left[\log Y | \mathbf{X}= \mathbf{x} \right] \right) \] cannot be interpreted with respect to original scale by simple \(\exp\) transformation \[ \mathsf{P} \left( (\exp\{LCI\}, \exp\{UCI\})\ni \exp\{\mathbf{x}^\top \widetilde{\boldsymbol{\beta}}\} = \exp\left\{\mathsf{E}\left[\log Y | \mathbf{X}= \mathbf{x} \right]\right\} < \mathsf{E}\left[Y | \mathbf{X}= \mathbf{x} \right] \right). \] However, it still works for prediction intervals: \[ \mathsf{P} \left( \log Y_{new} \in (LPI, UPI) \right) = \mathsf{P} \left( Y_{new} \in (\exp\{LPI\}, \exp\{UPI\}) \right). \] Now we plot the same outputs but on original scales. Grey colors correspond to incorrect estimates for \(\exp {\mathsf{E} [\log Y | \mathbf{X} ]}\) that are simple \(\exp\) transformation of the estimates for \(\mathsf{E} [\log Y | \mathbf{X} ]\). Using \(\Delta\)-method and knowledge of log-normal distribution, \(\mathsf{E} \exp{ \varepsilon} = \exp \left\{\frac{\sigma^2}{2}\right\}\), we can calculate the true estimates for \(\mathsf{E} \left[ Y | \mathbf{X}\right]\) (colourful).
YLIM <- range(chicago[, "fire"], exp(pfit))
XLIM <- range(chicago[, "minor"])
# True point estimate and CI for E[Y|X] based on log-N distribution
sigma2 <- summary(m)$sigma^2
EYX_fit <- exp(cifit$fit[,"fit"] + sigma2/2)
CIEYX_lwr <- EYX_fit * (1 - qnorm(0.975) * cifit$se.fit)
CIEYX_upr <- EYX_fit * (1 + qnorm(0.975) * cifit$se.fit)
par(mfrow = c(1, 2), mar=c(4,4,2,0.5))
for (ss in levels(chicago[, "fside"])){
plot(fire ~ minor, data = subset(chicago, fside == ss), main = ss,
xlab = XLAB, ylab = "Fires", xlim = XLIM, ylim = YLIM,
col = COLS[fside], bg = BGS[fside], pch = PCHS[fside])
ind <- if (ss == "North") 1:lpred else (lpred + 1):(2 * lpred)
lines(pminor, exp(cifit$fit[ind, "fit"]), col = "grey30", lwd = 3)
lines(pminor, exp(cifit$fit[ind, "lwr"]), col = "grey40", lwd = 2, lty = 2)
lines(pminor, exp(cifit$fit[ind, "upr"]), col = "grey40", lwd = 2, lty = 2)
lines(pminor, EYX_fit[ind], col = COLS[ss], lwd = 3)
lines(pminor, CIEYX_lwr[ind], col = COLS[ss], lwd = 2, lty = 2)
lines(pminor, CIEYX_upr[ind], col = COLS[ss], lwd = 2, lty = 2)
lines(pminor, exp(pfit[ind, "lwr"]), col = COLS[ss], lwd = 2, lty = 3)
lines(pminor, exp(pfit[ind, "upr"]), col = COLS[ss], lwd = 2, lty = 3)
legend("topleft", bty = "n",
legend=c("estim. exp(E[log(Y)|X=x]) < E[Y|X=x]",
"conf. band around exp(E[log(Y)|X=x]) < E[Y|X=x]",
"estim. E[Y|X=x]",
"conf. band around E[Y|X=x]",
"prediction band for Y"),
lwd=c(3,2,3,2,2), lty=c(1,2,1,2,3),
col=c("grey30","grey40",COLS[ss],COLS[ss],COLS[ss]))
}